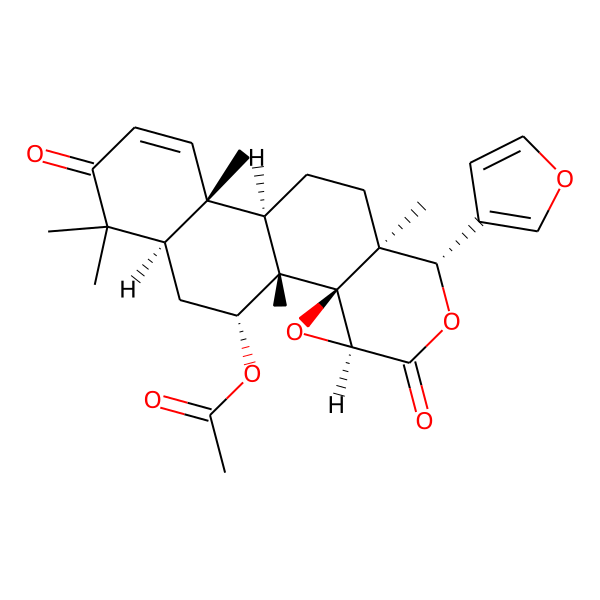
Gedunin
Function
Structures
Docking in target protein
Off-target analysis based on ligand similarity (Homo sapiens)
Step 1 - Target prediction for Gedunin: SwissTargetPrediction
Tips: Click on the link to jump to the 'SwissTargetPrediction' webserver. Select the species of 'Homo sapiens', and then paste the SMILES of Gedunin in the SMILES input box.
Step 2 - Blind docking for Gedunin: CB-Dock
Tips: Click on the link to jump to the 'CB-Dock' webserver. Upload the structure file of target predicted by 'SwissTargetPrediction' and the 2D/3D structure file of Gedunin to perform blind docking.