CB-Dock
Cavity-detection guided Blind Docking
CB-Dock is a protein-ligand docking method which automatically identifies the binding sites, calculates the center and size, customizes the docking box size according to the query ligands and then perform the molecular docking with AutoDock Vina. Large-scale benchmarks show that the cavity-focused docking can enhance the hit ratio and accuracy of blind docking. Accordingly, CB-Dock can facilitate the docking procedure and improve the accuracy by predicting the binding sites of target proteins using our curvature-based cavity detection approach (CurPocket) and the binding poses of query ligands using AutoDock Vina.
| Curvature of protein surface | Cavity detected by clustering | Docking with AutoDock Vina |
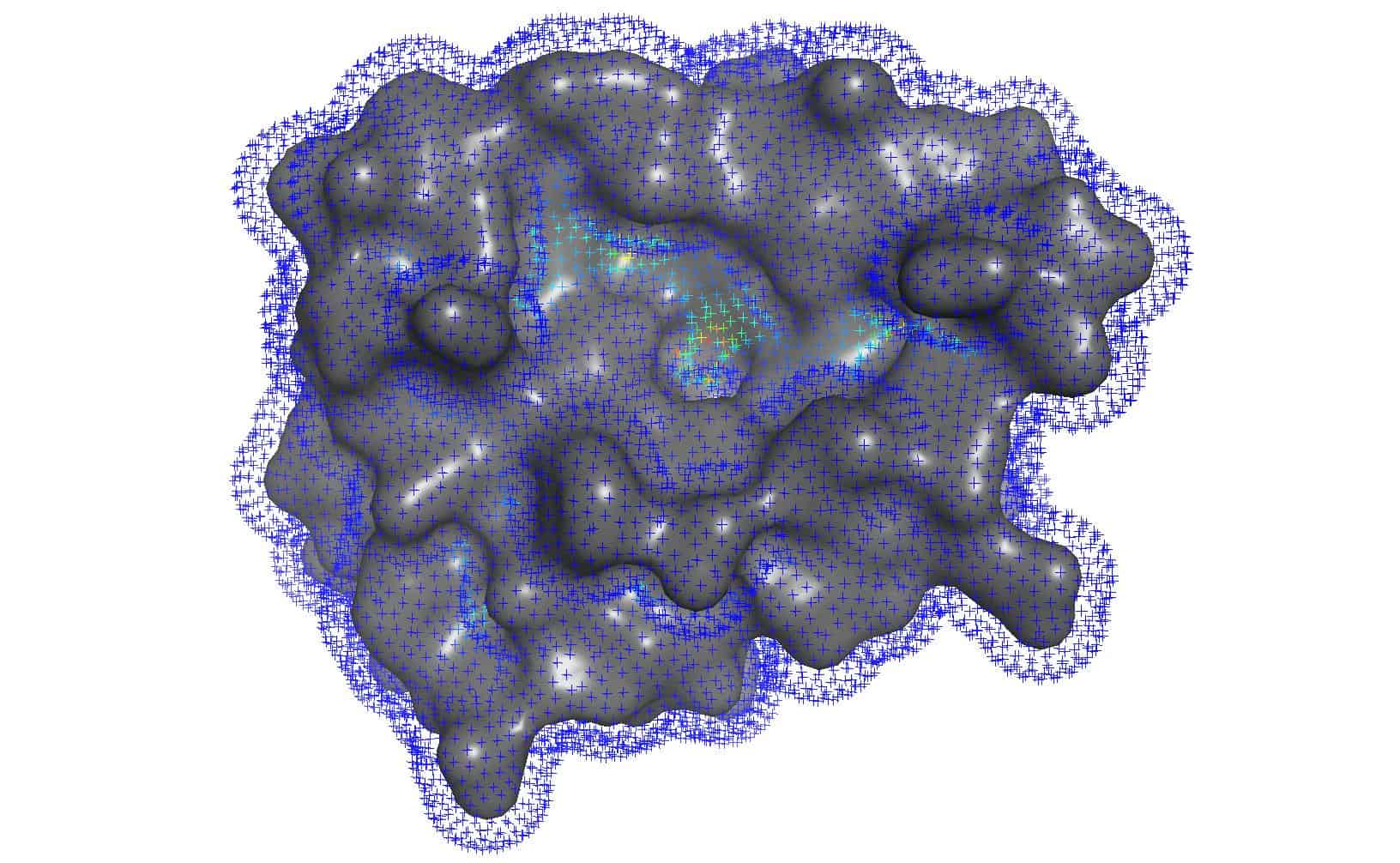
|
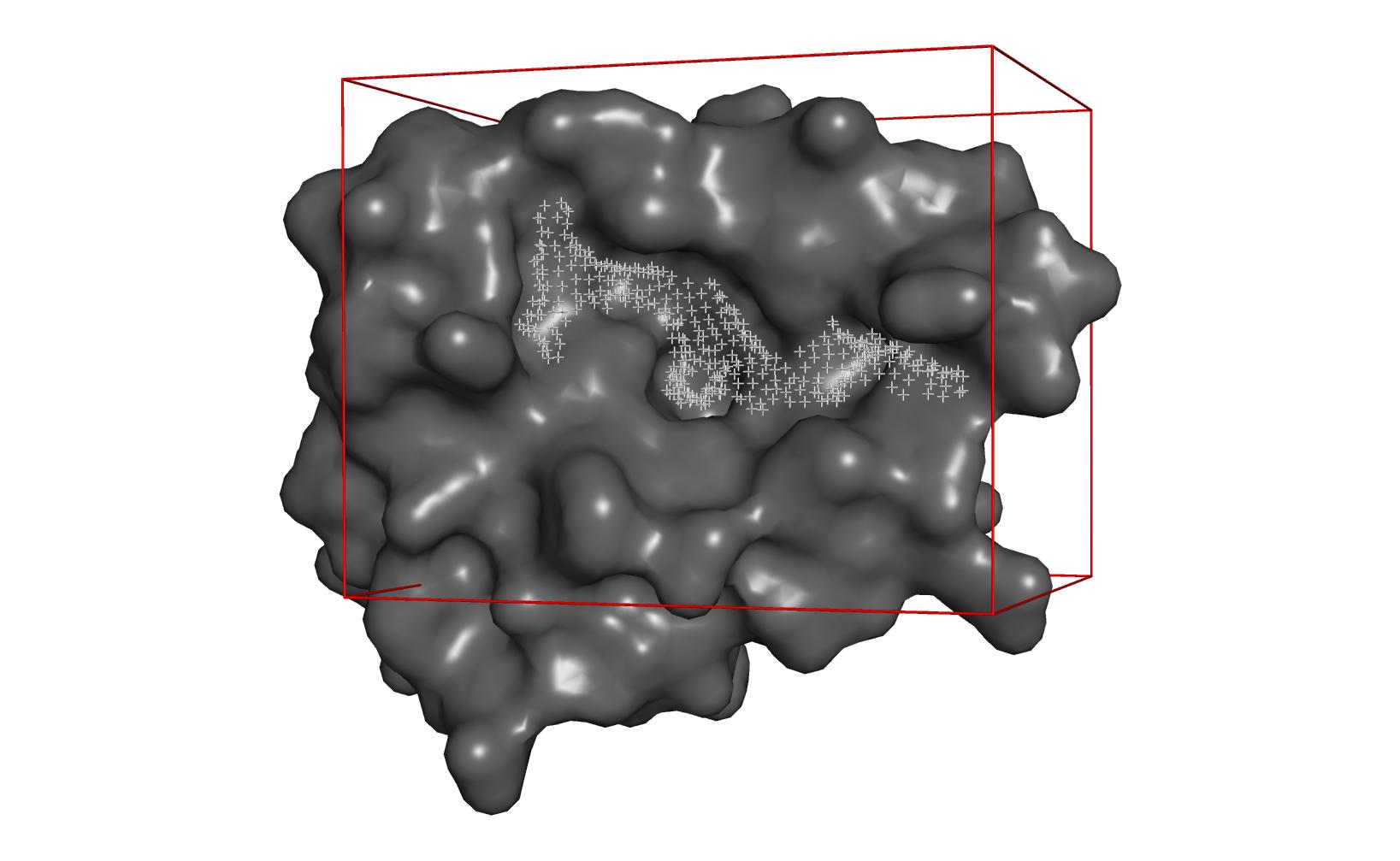

|
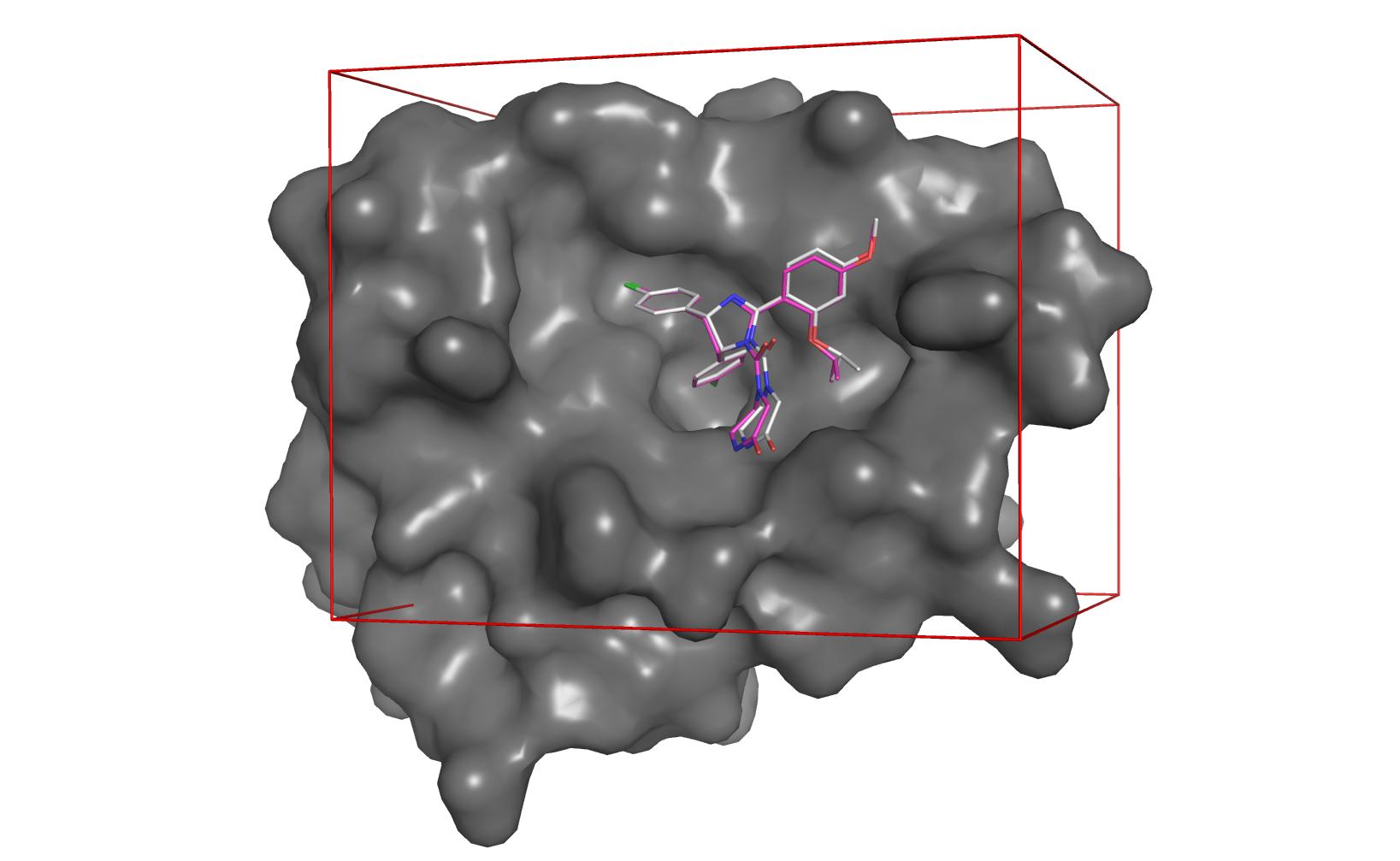
|
New Version Available
N
E
W
A new version of CB-Dock is available at http://cadd.labshare.cn/cb-dock2/.
Publication
Yang Liu, Maximilian Grimm, et al . CB-Dock: a web server for cavity detection-guided protein–ligand blind docking. Acta Pharmacologica Sinica, 2019.
Yang Cao , Lei Li. Improved protein-ligand binding affinity prediction by using a curvature dependent surface area model, . Bioinformatics, 2014.